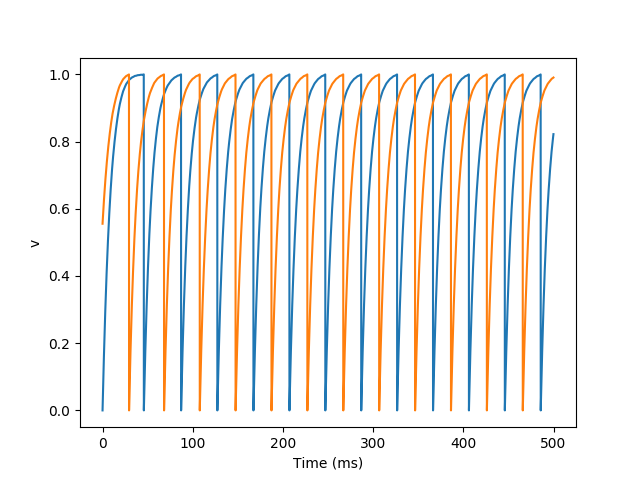
Example: gapjunctions
Neurons with gap junctions.
from brian2 import *
n = 10
v0 = 1.05
tau = 10*ms
eqs = '''
dv/dt = (v0 - v + Igap) / tau : 1
Igap : 1 # gap junction current
'''
neurons = NeuronGroup(n, eqs, threshold='v > 1', reset='v = 0',
method='exact')
neurons.v = 'i * 1.0 / (n-1)'
trace = StateMonitor(neurons, 'v', record=[0, 5])
S = Synapses(neurons, neurons, '''
w : 1 # gap junction conductance
Igap_post = w * (v_pre - v_post) : 1 (summed)
''')
S.connect()
S.w = .02
run(500*ms)
plot(trace.t/ms, trace[0].v)
plot(trace.t/ms, trace[5].v)
xlabel('Time (ms)')
ylabel('v')
show()