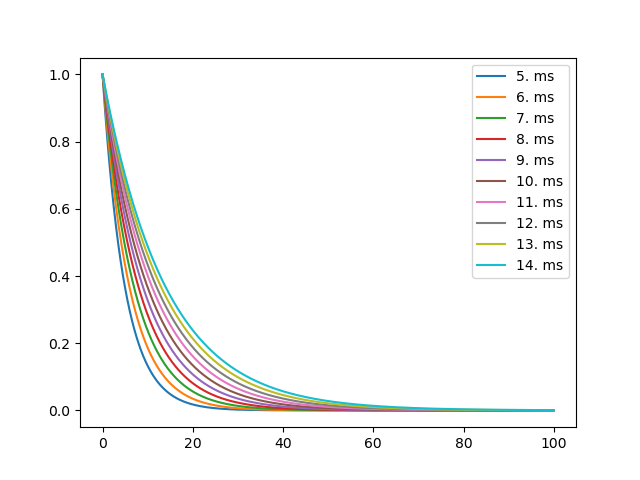
Example: 03_standalone_joblib
This example use C++ standalone mode for the simulation and the
joblib library
to parallelize the code. See the previous example (02_using_standalone.py)
for more explanations.
from joblib import Parallel, delayed
from time import time as wall_time
from brian2 import *
import os
def run_sim(tau):
pid = os.getpid()
directory = f"standalone{pid}"
set_device('cpp_standalone', directory=directory)
print(f'RUNNING {pid}')
G = NeuronGroup(1, 'dv/dt = -v/tau : 1', method='euler')
G.v = 1
mon = StateMonitor(G, 'v', record=0)
net = Network()
net.add(G, mon)
net.run(100 * ms)
res = (mon.t/ms, mon.v[0])
device.reinit()
print(f'FINISHED {pid}')
return res
if __name__ == "__main__":
start_time = wall_time()
n_jobs = 4
tau_values = np.arange(10)*ms + 5*ms
results = Parallel(n_jobs=n_jobs)(map(delayed(run_sim), tau_values))
print(f"Done in {wall_time() - start_time:10.3f}")
for tau_value, (t, v) in zip(tau_values, results):
plt.plot(t, v, label=str(tau_value))
plt.legend()
plt.show()