Example: standalone_multiplerun
This example shows how to run several, independent simulations in standalone mode.
Given that this example only involves a single neuron, an alternative – and arguably more elegant – solution
would be to run the simulations in a single NeuronGroup, where each neuron receives input with a different rate.
The example is a standalone equivalent of the one presented in /tutorials/3-intro-to-brian-simulations.
import numpy as np
import matplotlib.pyplot as plt
import brian2 as b2
from time import time
b2.set_device('cpp_standalone', build_on_run=False)
if __name__ == "__main__":
start_time = time()
num_inputs = 100
input_rate = 10 * b2.Hz
weight = 0.1
net = b2.Network()
P = b2.PoissonGroup(num_inputs, rates=input_rate)
eqs = """
dv/dt = -v/tau : 1
tau : second (constant)
"""
G = b2.NeuronGroup(1, eqs, threshold='v>1', reset='v=0', method='euler')
S = b2.Synapses(P, G, on_pre='v += weight')
S.connect()
M = b2.SpikeMonitor(G)
net.add([P, G, S, M])
net.run(1000 * b2.ms)
b2.device.build(run=False) # compile the code, but don't run it yet
npoints = 15
tau_range = np.linspace(1, 15, npoints) * b2.ms
output_rates = np.zeros(npoints)
for ii in range(npoints):
tau_i = tau_range[ii]
b2.device.run(run_args={G.tau: tau_i})
output_rates[ii] = M.num_spikes / b2.second
print(f"Done in {time() - start_time}")
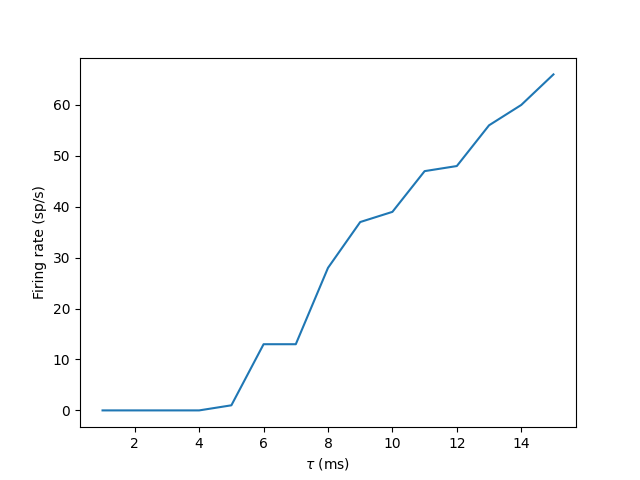
plt.plot(tau_range/b2.ms, output_rates)
plt.xlabel(r"$\tau$ (ms)")
plt.ylabel("Firing rate (sp/s)")
plt.show()