Example: Naud_et_al_2008_adex_firing_patterns
Firing patterns in the adaptive exponential integrate-and-fire model
Naud R et al. (2008): Firing patterns in the adaptive exponential integrate-and-fire model. Biol Cybern. 2008; 99(4): 335–347. doi:10.1007/s00422-008-0264-7
Parameters adapted by P. Müller to match figures, cf. http://www.kip.uni-heidelberg.de/Veroeffentlichungen/details.php?id=3445.
Sebastian Schmitt, Sebastian Billaudelle, 2022
from brian2 import *
import matplotlib.pyplot as plt
def sim(ax_vm, ax_w, ax_vm_w, parameters):
"""
simulate with parameters and plot to axes
"""
# taken from Touboul_Brette_2008
eqs = """
dvm/dt = (g_l*(e_l - vm) + g_l*d_t*exp((vm-v_t)/d_t) + i_stim - w)/c_m : volt
dw/dt = (a*(vm - e_l) - w)/tau_w : amp
"""
neuron = NeuronGroup(
1,
model=eqs,
threshold="vm > 0*mV",
reset="vm = v_r; w += b",
method="euler",
namespace=parameters,
)
neuron.vm = parameters["e_l"]
neuron.w = 0
states = StateMonitor(neuron, ["vm", "w"], record=True, when="thresholds")
defaultclock.dt = 0.1 * ms
run(0.6 * second)
# clip membrane voltages to threshold (0 mV)
vms = np.clip(states[0].vm / mV, a_min=None, a_max=0)
ax_vm.plot(states[0].t / ms, vms)
ax_w.plot(states[0].t / ms, states[0].w / nA)
ax_vm_w.plot(vms, states[0].w / nA)
ax_w.sharex(ax_vm)
ax_vm.tick_params(labelbottom=False)
ax_vm.set_ylabel("V [mV]")
ax_w.set_xlabel("t [ms]")
ax_w.set_ylabel("w [nA]")
ax_vm_w.set_xlabel("V [mV]")
ax_vm_w.set_ylabel("w [nA]")
ax_vm_w.yaxis.tick_right()
ax_vm_w.yaxis.set_label_position("right")
patterns = {
"tonic spiking": {
"c_m": 200 * pF,
"g_l": 10 * nS,
"e_l": -70.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": 2.0 * nS,
"tau_w": 30.0 * ms,
"b": 0.0 * pA,
"v_r": -58.0 * mV,
"i_stim": 500 * pA,
},
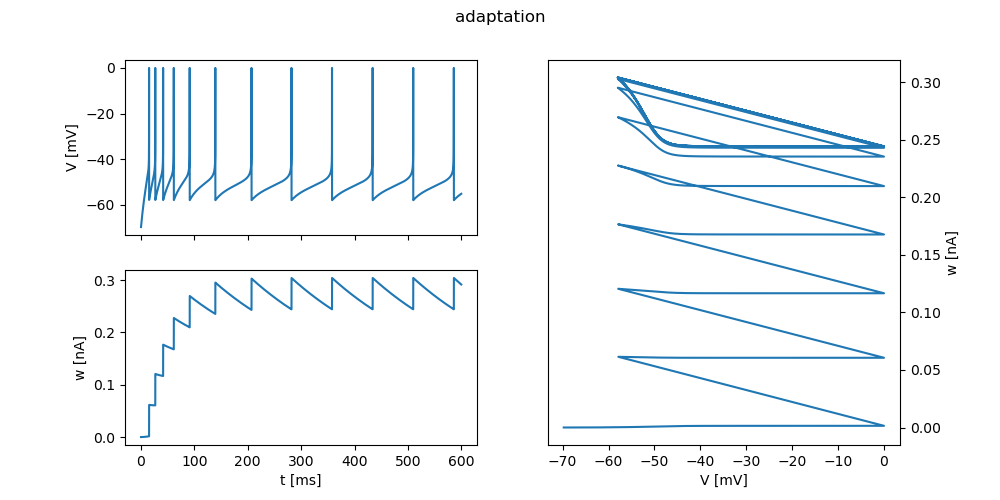
"adaptation": {
"c_m": 200 * pF,
"g_l": 12 * nS,
"e_l": -70.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": 2.0 * nS,
"tau_w": 300.0 * ms,
"b": 60.0 * pA,
"v_r": -58.0 * mV,
"i_stim": 500 * pA,
},
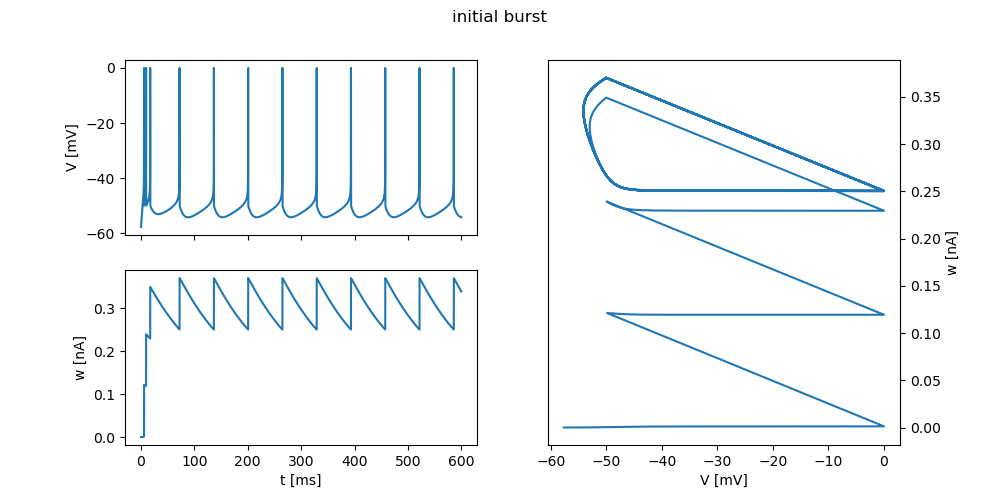
"initial burst": {
"c_m": 130 * pF,
"g_l": 18 * nS,
"e_l": -58.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": 4.0 * nS,
"tau_w": 150.0 * ms,
"b": 120.0 * pA,
"v_r": -50.0 * mV,
"i_stim": 400 * pA,
},
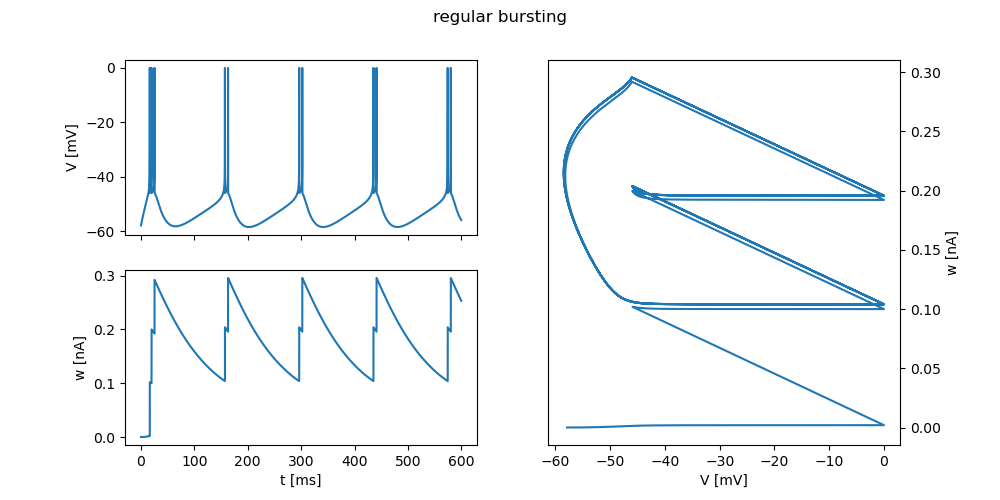
"regular bursting": {
"c_m": 200 * pF,
"g_l": 10 * nS,
"e_l": -58.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": 2.0 * nS,
"tau_w": 120.0 * ms,
"b": 100.0 * pA,
"v_r": -46.0 * mV,
"i_stim": 210 * pA,
},
"delayed accelerating": {
"c_m": 200 * pF,
"g_l": 12 * nS,
"e_l": -70.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": -10.0 * nS,
"tau_w": 300.0 * ms,
"b": 0.0 * pA,
"v_r": -58.0 * mV,
"i_stim": 300 * pA,
},
"delayed regular bursting": {
"c_m": 100 * pF,
"g_l": 10 * nS,
"e_l": -65.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": -10.0 * nS,
"tau_w": 90.0 * ms,
"b": 30.0 * pA,
"v_r": -47.0 * mV,
"i_stim": 110 * pA,
},
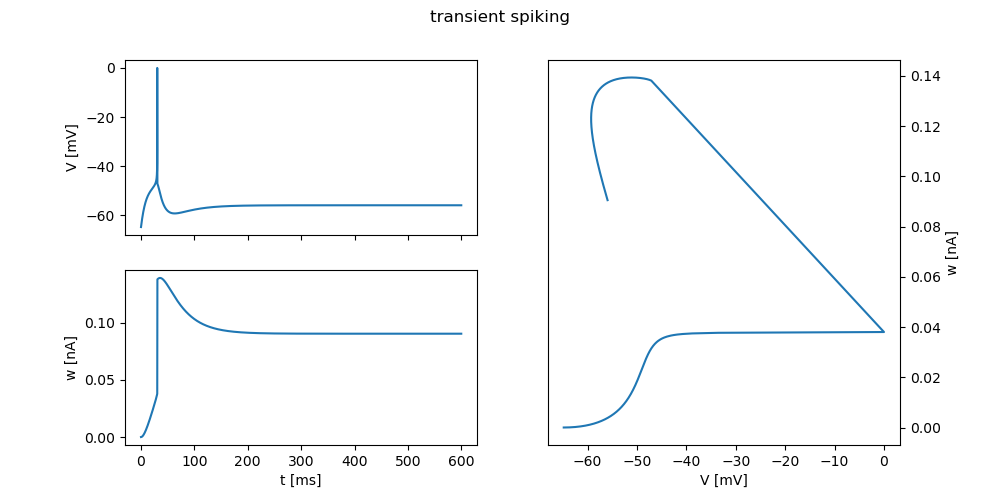
"transient spiking": {
"c_m": 100 * pF,
"g_l": 10 * nS,
"e_l": -65.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": 10.0 * nS,
"tau_w": 90.0 * ms,
"b": 100.0 * pA,
"v_r": -47.0 * mV,
"i_stim": 180 * pA,
},
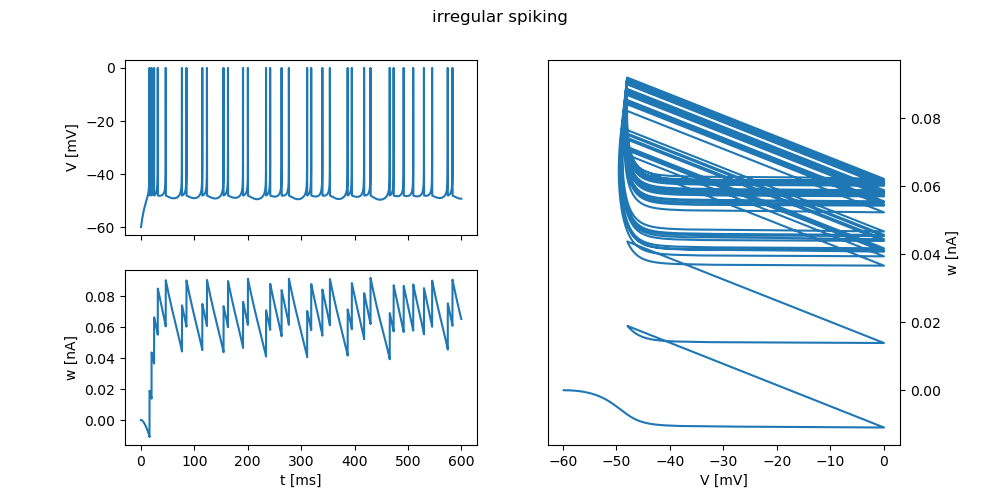
"irregular spiking": {
"c_m": 100 * pF,
"g_l": 12 * nS,
"e_l": -60.0 * mV,
"v_t": -50.0 * mV,
"d_t": 2.0 * mV,
"a": -11.0 * nS,
"tau_w": 130.0 * ms,
"b": 30.0 * pA,
"v_r": -48.0 * mV,
"i_stim": 160 * pA,
},
}
# loop over all patterns and plot
for pattern, parameters in patterns.items():
fig = plt.figure(figsize=(10, 5))
fig.suptitle(pattern)
gs = fig.add_gridspec(2, 2)
ax_vm = fig.add_subplot(gs[0, 0])
ax_w = fig.add_subplot(gs[1, 0])
ax_vm_w = fig.add_subplot(gs[:, 1])
sim(ax_vm, ax_w, ax_vm_w, parameters)
plt.show()