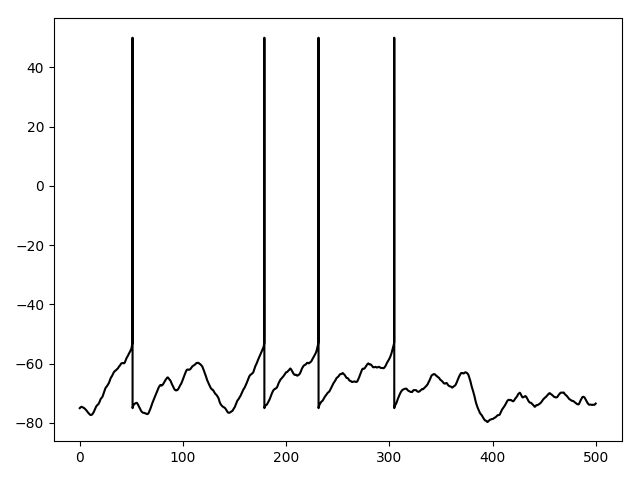
Example: Fig5A
Brette R (2013). Sharpness of spike initiation in neurons explained by compartmentalization. PLoS Comp Biol, doi: 10.1371/journal.pcbi.1003338.
Fig. 5A. Voltage trace for current injection, with an additional reset when a spike is produced.
Trick: to reset the entire neuron, we use a set of synapses from the spike initiation compartment where the threshold condition applies to all compartments, and the reset operation (v = EL) is applied there every time a spike is produced.
from brian2 import *
from params import *
defaultclock.dt = 0.025*ms
duration = 500*ms
# Morphology
morpho = Soma(50*um) # chosen for a target Rm
morpho.axon = Cylinder(diameter=1*um, length=300*um, n=300)
# Input
taux = 5*ms
sigmax = 12*mV
xx0 = 7*mV
compartment = 40
# Channels
eqs = '''
Im = gL * (EL - v) + gNa * m * (ENa - v) + gLx * (xx0 + xx) : amp/meter**2
dm/dt = (minf - m) / taum : 1 # simplified Na channel
minf = 1 / (1 + exp((va - v) / ka)) : 1
gNa : siemens/meter**2
gLx : siemens/meter**2
dxx/dt = -xx / taux + sigmax * (2 / taux)**.5 *xi : volt
'''
neuron = SpatialNeuron(morphology=morpho, model=eqs, Cm=Cm, Ri=Ri,
threshold='m>0.5', threshold_location=compartment,
refractory=5*ms)
neuron.v = EL
neuron.gLx[0] = gL
neuron.gNa[compartment] = gNa_0 / neuron.area[compartment]
# Reset the entire neuron when there is a spike
reset = Synapses(neuron, neuron, on_pre='v = EL')
reset.connect('i == compartment') # Connects the spike initiation compartment to all compartments
# Monitors
S = SpikeMonitor(neuron)
M = StateMonitor(neuron, 'v', record=0)
run(duration, report='text')
# Add spikes for display
v = M[0].v
for t in S.t:
v[int(t / defaultclock.dt)] = 50*mV
plot(M.t/ms, v/mV, 'k')
tight_layout()
show()